ERP Core N400 in MNE-Python: part III
In the last post on adapting ERP CORE’s N400 pipeline to MNE-Python, we did the ICA-related steps to correct blinking and horizontal eye-movement artfacts from the signal. Here we do some final steps in cleaning the data: interpolation of bad channels, epoching and rejection of epochs with possible artifacts. These steps are decipted in the Elist_Bin_Epoch script of ERP CORE resources (script 5).
Contrary to previous posts, I won’t make the whole process in a single participant and show the loop for all participants at the end. That is because the result for some af the steps are saved as intermediate stages of the preprocessing pipeline. I should also tell you up-front that I wasn’t able to find implemented equivalents for some of the procedures ERP CORE uses to define artifacts. That forced me to implement a python version of what they do. Therefore, this post is a little longer the the other ones, so buckle up! On the bright side, at the end we will have the data ready to start looking for those N400 ERPs.
Interpolation
Our first step here is to interpolate bad channels. Bad channels for each participant are listed in the file “Interpolate_Channels_N400.xlsx” provided by ERP CORE. Below, I load the information in that file, make a list of bad channels for each participant and add that list to their raw.info['bads'].
Load the information:
import os
os.chdir('D:\\EEGdata\\Erp CORE\\N400')
import mne
# import some methods directly so we can call them by name
from mne.io import read_raw
from mne.epochs import read_epochs
import numpy as np
import pandas as pd
%matplotlib qt
%gui qt
# load as DataFrame
interpolate_path = "D:\\EEGdata\\Erp CORE\\N400\\EEG_ERP_Processing\\Interpolate_Channels_N400.xlsx"
chans_interpolate = pd.read_excel(interpolate_path)
# Turn strings to a list of strings for each participant
chans_interpolate = chans_interpolate.loc[:,"Name of Interpolated Channel"].str.split(" ")
Now I’ll deal with some nuisences in the file: there is an asterisk at the beginning of the channel list for each participant, and Fp channels are in upper-case. We remove the asterisk and correct the captalization.
# make function to remove asterisk and correct
def remove_star(item):
# if item is a list
if isinstance(item, list):
# if list is longer than 1
if len(item) > 1:
# remove asterisk
clean_names = [item[0].lstrip("*")]
# add channel to a list clean_names
clean_names.extend(item[1:])
# look for Fp channels and correct captalization
for n, elec in enumerate(clean_names):
if elec =='FP1' or elec =='FP2':
clean_names[n] = elec.capitalize()
else:
clean_names = [item[0].lstrip("*")]
if clean_names[0] =='FP1' or clean_names[0] =='FP2':
clean_names[0] = clean_names[0].capitalize()
else:
clean_names = np.nan
return clean_names
# apply function
chans_interpolate = chans_interpolate.apply(remove_star)
chans_interpolate.loc[chans_interpolate.notnull()].head()
0 [F8]
4 [Fp2]
5 [FC4, PO4]
10 [P7]
16 [F8]
Name: Name of Interpolated Channel, dtype: object
We have already indentified the channels to be interpolated, however if we try to do the interpolation now MNE will throw us an error because we have more EEG channels than mapped on the 3D digitation. The channels HEOG_left, HEOG_right and VEOG_lower are currently identified as EEG channels, but they are actually extra channels we used to make the proper HEOG and VEOG channels. We will drop them, since they already served their purpose.
Let’s do it for all participants and save an intermediate (“interpol”) raw file:
for subj in range(1,41):
raw_correct_path = "D:\\EEGdata\\Erp CORE\\N400\\mne\\raw_corrected\\N400_ERP_CORE_{}_correct-raw.fif".format(subj)
raw_correct = read_raw(raw_correct_path, preload = True)
#only do interpolation if there is at least one channel to interpolate
if not chans_interpolate.isna()[subj-1]:
raw_correct.info['bads'].extend(chans_interpolate[subj-1])
# drop extra channels
raw_correct.drop_channels(['HEOG_left', 'HEOG_right', 'VEOG_lower'])
raw_correct.interpolate_bads()
else:
raw_correct.drop_channels(['HEOG_left', 'HEOG_right', 'VEOG_lower'])
interpol_path = "D:\\EEGdata\\Erp CORE\\N400\\mne\\raw_interpol\\N400_ERP_CORE_{}_interpol-raw.fif".format(subj)
raw_correct.save(interpol_path, overwrite=True)
Event list for epochs
Now let’s make our event list and event_ids for epoching.
As we did at the back at beginning of preprocessing
(in this post), we will get the events from the annotations in the data. Until now, we have two types of annotation descriptions in the data: “BAD_seg” and numbers corresponding to triggers used in the experiment. The ERP CORE file “N400_Event_Code_Scheme.xlsx” has the meaning for each code. For instance, the code 121 refers to a Priming stimuli which is unrelated to the target stimulus and comes from the second list of words; while the code 201 means that the participant gave a correct answer.
We are going to change these codes to a descriptive account of each event. A nice feature of MNE is that we can make hierarchical ids. We will follow the hierarchy shown in the N400_Event_Code_Scheme file: Word Type > Relatedness > Word List.
First, we will create a dictionary with the correspondence between codes and events to properly identify them later. We will be able to use these ids to select epochs afterwards.
# load data
subj = 1
interpol_path = "D:\\EEGdata\\Erp CORE\\N400\\mne\\raw_interpol\\N400_ERP_CORE_{}_interpol-raw.fif".format(subj)
raw_interpol = read_raw(interpol_path)
# make dict with correspondences between codes and hierarchical ids
description_dict = {'111' : 'Prime/Related/L1',
'112' : 'Prime/Related/L2',
'121' : 'Prime/Unrelated/L1',
'122' : 'Prime/Unrelated/L2',
'211' : 'Target/Related/L1',
'212' : 'Target/Related/L2',
'221' : 'Target/Unrelated/L1',
'222' : 'Target/Unrelated/L2',
'201' : 'Hit',
'202' : 'Miss',
'BAD_seg': 'BAD_seg'
}
We will change the description list with some Pandas functionality. It is beyond this tutorial to teach Pandas’ skills, but it is nice for you to know that, although most of the backbone of MNE is in Numpy, it also allows exportation to Pandas’ data structures.
In summary, what we do in the one-liner below is make a copy of the annotations descriptions while transforming it to a Pandas series, then we make the substituitions according to our description dictionary and turn it back to a numpy array.
raw_interpol.annotations.description = pd.Series(raw_interpol.annotations.description).map(description_dict).to_numpy()
# substitute annotations description
set(raw_interpol.annotations.description)
Now we are almost ready to make the events numpy array we will need to epoch the data. We will use the events_from_annotations function. One last step before that: create a dictionary with correspondences between hierarchical ids and trigger numbers.
You may be wondering why we would do that when we just made the reverse dictionary on our last step. The thing is that MNE’s event list is a numeric array composed of columns instant, blank column and event code, and the default behavior of events_from_annotations is to assign the event codes as [0,1,2,3…]. However, I want to keep the original codes from ERP CORE, so I need to use the events_id input with a dict showing the correspondences. I should say that for the rest of our MNE operations the numeric codes themselves are actually rather irrelevant, what really matters are the correspondences in event_ids. I am just being picky to keep as close as possible to the original ERP CORE material.
event_ids = {'Prime/Related/L1': 111,
'Prime/Related/L2': 112,
'Prime/Unrelated/L1' : 121,
'Prime/Unrelated/L2' : 122,
'Target/Related/L1' : 211,
'Target/Related/L2' : 212,
'Target/Unrelated/L1' : 221,
'Target/Unrelated/L2' : 222,
'Hit' : 201,
'Miss' : 202}
events, event_ids = mne.events_from_annotations(raw_interpol, event_ids)
So, our events look like this (instant, blank column, event code):
events
array([[ 3058, 0, 202],
[ 4145, 0, 202],
[ 5195, 0, 112],
...,
[144827, 0, 121],
[145156, 0, 221],
[145325, 0, 201]])
Epoching and artfact rejection
The ERP CORE pipeline is very throrough in its epoch dropping due to ocular artifact detection. There are different thresholds for epoch dropping for: (1) EEG channels, (2) the VEOG channel and (3 and 4) for both variations of the HEOG channel (with and without ICA applied). In MNE, we can set epoch rejection thresholds for each channel type by setting the reject input when we create the epochs.
However, as far as I understand from the documentation, MNE uses a simple peak-to-peak algorithm to do its automatic rejection. ERP CORE pipeline uses such an algorithm only for rejection on the scalp EEG channels. For VEOG and uncorrected HEOG channels it uses a windowed peak-to-peak algorithm and for ICA corrected HEOG it uses a step algorithm (both as implemented in the ERP LAB toolbox).
Since ERP CORE gives us the parameters, and the algorithms themselves are quite straightforward, I have implemented them below. I make use of the nifty feature of MNE that allows us to loop through epochs. On each epoch I run the algorithms on the corresponding channels and, if some amplitude is above the threshold, it is marked as bad with a MW_Reject annotation. The value compared to the treshold on each algorithm are the following:
- Step algorithm: $\bmod (mean(window1)-mean(window2))$
- Peak-to-peak algorithm: $\bmod (max(window1)-min(window2))$
Below, we (1) load the the information we need for the artfact rejection procedure, (2) create the dictionaries we need for marking the events in the experiment (as shown above); then, in the loop, we (3) add the annotations for epoching, (4) do the epoching with mne.Epochs(), (5) run the artfact rejection algorithm and (6) save the epochs.
#get eog channels numbers
subj = 1
interpol_path = "D:\\EEGdata\\Erp CORE\\N400\\mne\\raw_interpol\\N400_ERP_CORE_{}_interpol-raw.fif".format(subj)
raw_interpol = read_raw(interpol_path)
eog_nums = [np.where(np.array(raw_interpol.ch_names) == 'VEOG')[0].item(0),
np.where(np.array(raw_interpol.ch_names) == 'HEOG')[0].item(0),
np.where(np.array(raw_interpol.ch_names) == 'HEOG_ICA')[0].item(0)]
#get veog moving windows parameters from files
veog_path = "D:\\EEGdata\\Erp CORE\\N400\\EEG_ERP_Processing\\AR_Parameters_for_MW_CRAP_N400.xlsx"
veog_mw_params = pd.read_excel(veog_path)
#convert time columns to seconds
veog_mw_params.iloc[:,3:] = veog_mw_params.iloc[:,3:].apply(lambda x: x*1e-3)
#convert threshold to volts
veog_mw_params.Threshold = veog_mw_params.Threshold*1e-4
# the same for non ICA corrected heog moving windows
heog_path = "D:\\EEGdata\\Erp CORE\\N400\\EEG_ERP_Processing\\AR_Parameters_for_MW_Blinks_N400.xlsx"
heog_mw_params = pd.read_excel(heog_path)
heog_mw_params.iloc[:,3:] = heog_mw_params.iloc[:,3:].apply(lambda x: x*1e-3)
heog_mw_params.Threshold = heog_mw_params.Threshold*1e-4
#... and for ICA corrected HEOG moving windows
heog_ica_path = "D:\\EEGdata\\Erp CORE\\N400\\EEG_ERP_Processing\\AR_Parameters_for_SL_HEOG_N400.xlsx"
heog_ica_mw_params = pd.read_excel(heog_ica_path)
heog_ica_mw_params.iloc[:,3:] = heog_ica_mw_params.iloc[:,3:].apply(lambda x: x*1e-3)
heog_ica_mw_params.Threshold = heog_ica_mw_params.Threshold*1e-4
#put moving windows dfs in a list
mw_params = [veog_mw_params, heog_mw_params, heog_ica_mw_params]
# make dict with correspondences between codes and hierarchical ids
description_dict = {'111' : 'Prime/Related/L1',
'112' : 'Prime/Related/L2',
'121' : 'Prime/Unrelated/L1',
'122' : 'Prime/Unrelated/L2',
'211' : 'Target/Related/L1',
'212' : 'Target/Related/L2',
'221' : 'Target/Unrelated/L1',
'222' : 'Target/Unrelated/L2',
'201' : 'Hit',
'202' : 'Miss',
'BAD_seg': 'BAD_seg'
}
#create correspondence between description and trigger number
event_ids = {'Prime/Related/L1': 111,
'Prime/Related/L2': 112,
'Prime/Unrelated/L1' : 121,
'Prime/Unrelated/L2' : 122,
'Target/Related/L1' : 211,
'Target/Related/L2' : 212,
'Target/Unrelated/L1' : 221,
'Target/Unrelated/L2' : 222,
'Hit' : 201,
'Miss' : 202}
# loop
for subj in range(1,41):
interpol_path = "D:\\EEGdata\\Erp CORE\\N400\\mne\\raw_interpol\\N400_ERP_CORE_{}_interpol-raw.fif".format(subj)
raw_interpol = read_raw(interpol_path)
#add event descriptions to annotations
raw_interpol.annotations.description = pd.Series(raw_interpol.annotations.description).map(description_dict).to_numpy()
set(raw_interpol.annotations.description)
#create events and event ids
events, event_ids = mne.events_from_annotations(raw_interpol, event_ids)
#change heog channel types
raw_interpol.set_channel_types({'HEOG_ICA':'emg', 'HEOG':'misc'})
epochs = mne.Epochs(raw_interpol,
events,
event_ids,
baseline = (-.2,0),
tmax = .8,
reject = dict(eeg = 200e-6)
)
epos_to_drop = []
# run through every epoch
for i in range(len(epochs.events)):
# for each channel of interest
for eog_chan_num, mw_df in zip(eog_nums, mw_params): #
# get moving windows parameters
w_size = mw_df['Window Size'].iloc[subj-1]
w_start = mw_df['Time Window Minimum'].iloc[subj-1].round(2)
w_start_last = np.round(mw_df['Time Window Maximum'].iloc[subj-1] - mw_df['Window Size'].iloc[subj-1],2)
w_stop = mw_df['Time Window Maximum'].iloc[subj-1].round(2)
w_stop_first = np.round(w_start + w_size,2)
# create an array with starting and stopping times of every window
mw_starts = np.arange(w_start,
w_start_last,
w_size)
mw_stops = np.arange(w_stop_first,
w_stop,
w_size)
# only run if channel is not empty, e.g. already marked as bad
if epochs[i].get_data()[:,eog_chan_num,:].shape[0] != 0:
#get channel data
chan_data = epochs[i].get_data()[:,eog_chan_num,:].flatten()
#make list with data for each window
mws = [ chan_data[epochs.time_as_index(start).item(0) : epochs.time_as_index(stop).item(0)] for start, stop in zip(mw_starts, mw_stops)]
# if channel is veog or uncorrected heog do peak-to-peak artfact detection
if eog_chan_num < 32:
mws = pd.DataFrame(mws).T
peak2peak = mws.apply(lambda x: np.abs(x.max() - x.min()))
if any(peak2peak > mw_df['Threshold'].iloc[subj-1]):
epos_to_drop.append(i)
# if channel is heog_ica do step artfact detection
else:
steps = [True for i in range(len(mws)-1) if np.abs(mws[i].mean()-mws[i+1].mean()) > mw_df['Threshold'].iloc[subj-1]]
if steps:
epos_to_drop.append(i)
# if any epoch was set as bad add it to drop list with label MW Reject
if epos_to_drop:
epochs.drop(epos_to_drop,'MW_Reject')
#Save epochs
epochs_path = "D:\\EEGdata\\Erp CORE\\N400\\mne\\epochs\\N400_ERP_CORE_{}-epo.fif".format(subj)
print('Subject {}'.format(subj)) # line to keep up with analysis as it runs
epochs.save(epochs_path, overwrite=True)
Lastly, let’s check the percentage of dropped epochs for each participant.
percent_dropped = pd.DataFrame(data = {'Subj': np.repeat(np.nan, 40),
'Dropped': np.repeat(np.nan, 40)})
for subj in range(1,41):
epochs_path = "D:\\EEGdata\\Erp CORE\\N400\\mne\\epochs\\N400_ERP_CORE_{}-epo.fif".format(subj)
epochs = read_epochs(epochs_path)
percent_dropped['Subj'].iloc[subj-1] = subj
percent_dropped['Dropped'].iloc[subj-1] = np.round(epochs.drop_log_stats(),2)
percent_dropped['Dropped'].describe().round(2)
count 40.00
mean 13.14
std 18.53
min 3.88
25% 4.97
50% 5.80
75% 9.51
max 85.64
Name: Dropped, dtype: float64
Dropped epochs per participant range from roughly 4% to 85%. Here, we will set consider participants with more than 30% of epochs dropped as participants with too many artfacts to be used. The ones that do not pass this filter are below:
bad_subjs = percent_dropped.loc[percent_dropped.Dropped >30].Subj
bad_subjs.to_csv('bad_subjs.csv')
bad_subjs.count()
5
We have 5 unsuable participants. This is different from ERP CORE’s original preprocessing results, where they lose a single participant. However, I wasn’t able to exactly reproduce the ICA step in the pipeline, so that is probably where we are falling a little behind. Anyway, 35 participants are more than enough for us to grasp the N400 effects in the following analyses. So, enough preprocesing, let’s get to those ERPs in the next post!
Let’s finish contemplating some hard earned epochs:
epochs.plot(n_epochs = 4)
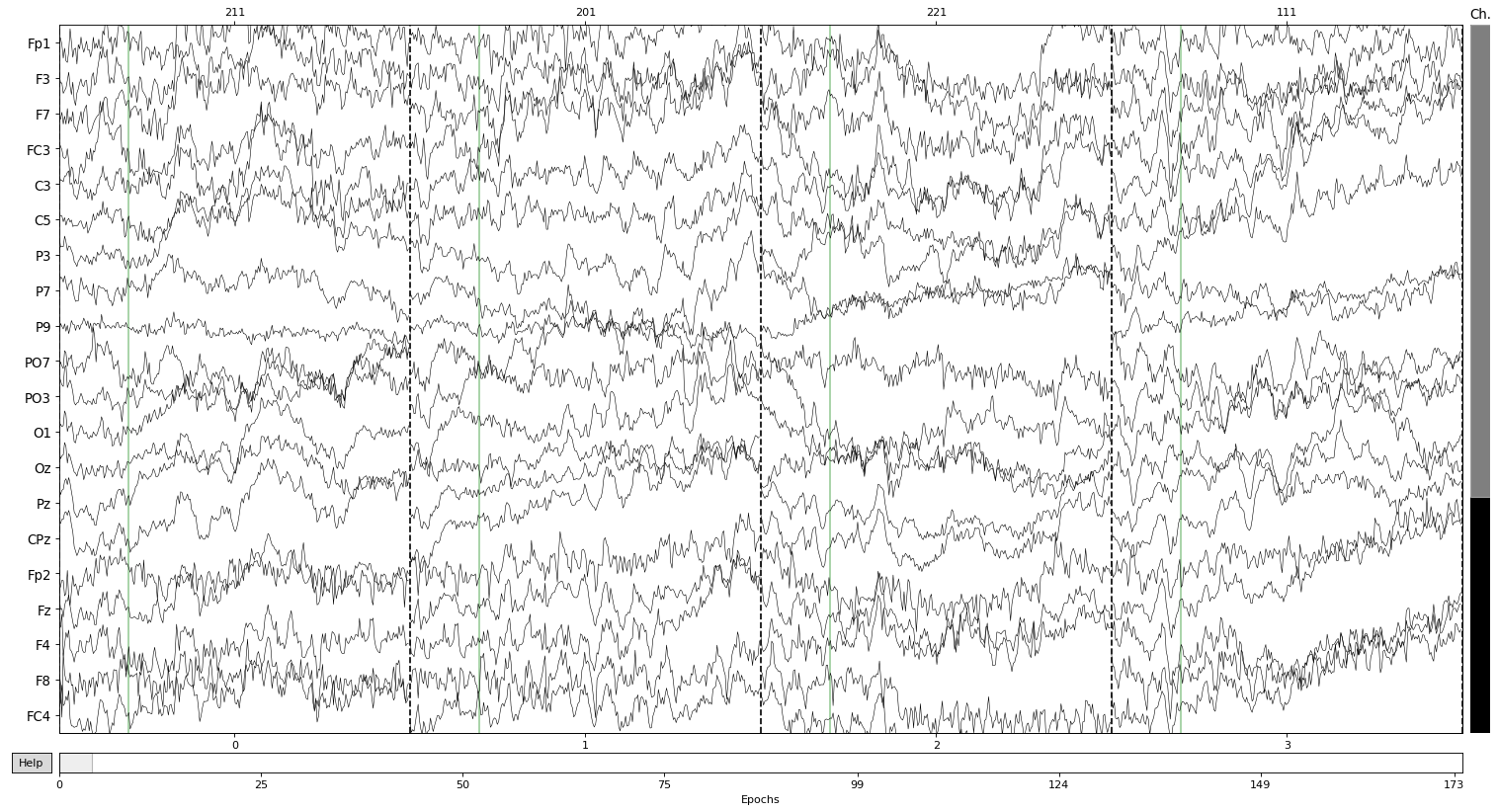
Dropped 0 epochs:
Channels marked as bad: []